Carmen Catalá
Assistant Professor
Investigating how plant hormones and cellular processes regulate fruit development, especially in tomato.

What are the processes involved in fruit formation?
Email: cc283@cornell.edu
Office/Lab: 315/316
Senior Research Associate
Section of Plant Biology
School of Integrative Plant Science
Cornell University
Philippe Nicolas, Richard J Pattison, Yi Zheng, Taly Lapidot-Cohen, Yariv Brotman, Sonia Osorio, Alisdair R Fernie, Zhangjun Fei, Carmen Catalá, Starch deficiency in tomato causes transcriptional reprogramming that modulates fruit development, metabolism, and stress responses, Journal of Experimental Botany, Volume 74, Issue 20, 31 October 2023, Pages 6331–6348
Philippe Nicolas, Yoshihito Shinozaki, Adrian Powell, Glenn Philippe, Stephen I Snyder, Kan Bao, Yi Zheng, Yimin Xu, Lance Courtney, Julia Vrebalov, Clare L Casteel, Lukas A Mueller, Zhangjun Fei, James J Giovannoni, Jocelyn K C Rose, Carmen Catalá, Spatiotemporal dynamics of the tomato fruit transcriptome under prolonged water stress, Plant Physiology, Volume 190, Issue 4, December 2022, Pages 2557–2578
Wang, X., Gao, L., Jiao, C. et al. Genome of Solanum pimpinellifolium provides insights into structural variants during tomato breeding. Nat Commun 11, 5817 (2020).
Shinozaki, Y., Nicolas, P., Fernandez-Pozo, N. et al. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening. Nat Commun 9, 364 (2018).
Noe Fernandez-Pozo, Yi Zheng, Stephen I Snyder, Philippe Nicolas, Yoshihito Shinozaki, Zhangjun Fei, Carmen Catala, James J Giovannoni, Jocelyn K.C Rose, Lukas A Mueller, The Tomato Expression Atlas, Bioinformatics, Volume 33, Issue 15, August 2017, Pages 2397–2398
Research Overview
Our research focuses on the molecular regulation of fruit development using tomato as a model system. Fruit development is a crucial process in the sexual reproduction of flowering plants and of critical importance for seed dispersal, plant fitness and agricultural yield. We use molecular and genetic techniques to investigate the complex interplay of gene expression changes, signaling events, and hormonal activity, controlling fruit development.
The tissue-specific transcript landscape of the developing tomato fruit
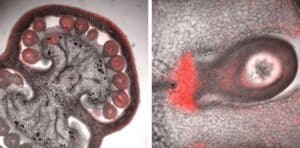
Fruit are complex organs, comprising distinct tissues and cell types, that arise from the coordinated growth and development of floral tissues following pollination and fertilization. We have used Laser Capture Microdissection (LCM) coupled with mRNA profiling (RNAseq) to analyze transcriptional changes during tomato fruit ontogeny with a high level of temporal and spatial resolution. We are building upon the data generated from our tissue-specific transcriptome studies by selecting candidate genes with potentially novel functions in the control of fruit development. (Pattison et al., 2015 https://doi.org/10.1104/pp.15.00287, Shinozaki et al., 2018 https://doi.org/10.1038/s41467-017-02782-9).
Fruit development under adverse environmental conditions
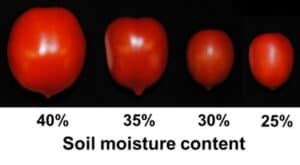
Successful fruit development after fertilization involves tightly regulated changes in gene expression, that can be severely affected by environmental stresses such as drought. We have analyzed the tissue specific transcriptional responses of tomato fruit developing under water deficit, and uncovered transcriptional networks controlling the responses of the fruit to abiotic stress (Nicolas et al., 2022 https://doi.org/10.1093/plphys/kiac445).
We are also investigating the role that sugars and starch metabolism play in regulating tissue or cell-specific responses to drought stress (Nicolas et al., 2023 https://doi.org/10.1093/jxb/erad212). The long-term goal of this research is to elucidate the molecular pathways underlying adaptations and responses to drought during to the plant reproductive phase.
One of our projects explores the genetic diversity offered by tomato wild species, which are adapted to extreme habitats, to reveal genes and pathways underlying successful fruit development under adverse environments. We are generating comprehensive tissue-specific transcriptomes of tomato wild relatives and examining regulatory variation using allele-specific expression analysis in hybrids with cultivated varieties. Our main goal is to increase our understanding of the molecular basis for phenotypic variation in fruit development.

Lab Members


Chiara Giraldo
In the News
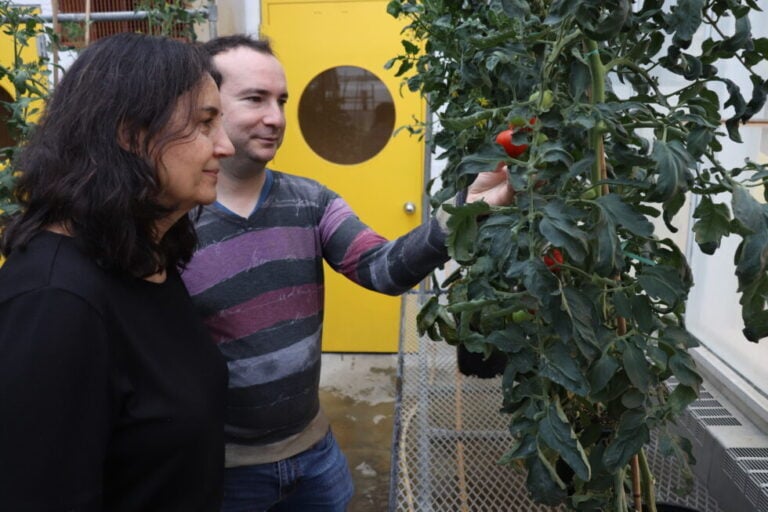

From Ground-Hugging to Groundbreaking: How a Unique Tomato Mutation Could Transform Sustainable Agriculture
omatoes are a staple in diets worldwide and an essential part of sustainable agriculture. Now, scientists at BTI have reported groundbreaking insights into a long-known tomato mutation, unlocking the potential...
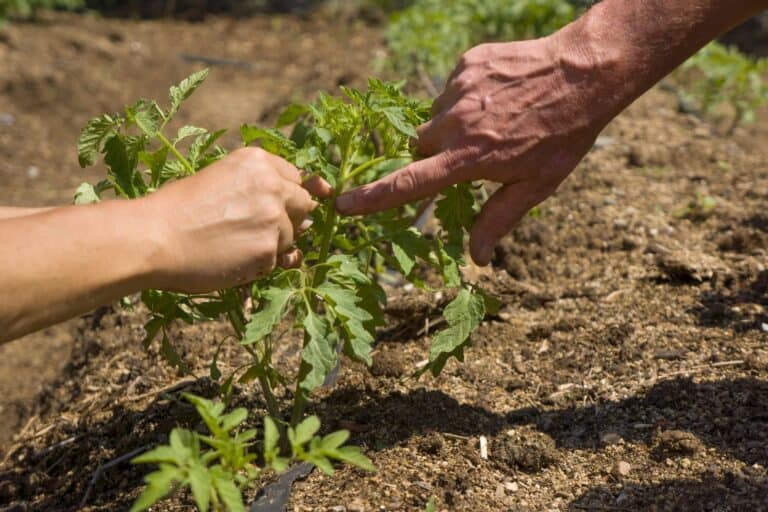

Finding genes to help fruit adapt to droughts
As shifting weather patterns are expected to lead to more frequent periods of drought, researchers are increasingly working to make discoveries that can help plants adapt to prolonged water stress....
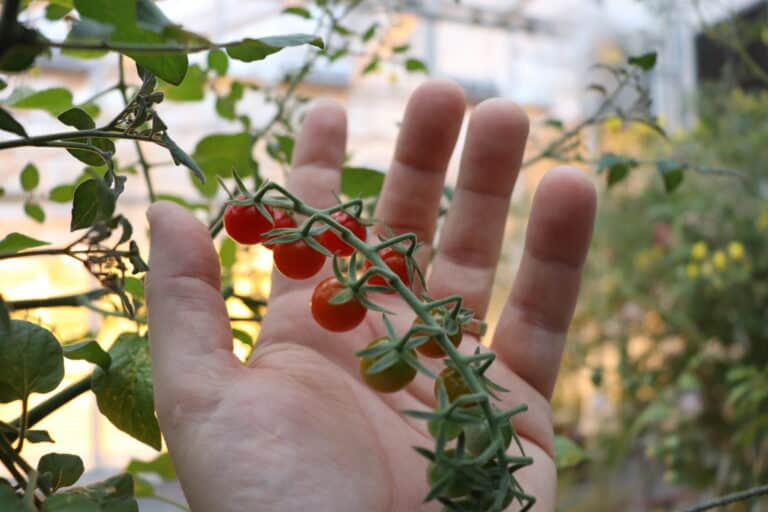

Tomato’s Wild Ancestor Is a Genomic Reservoir for Plant Breeders
Thousands of years ago, people in the region now known as South America began domesticating Solanum pimpinellifolium, a weedy plant with small, intensely flavored fruit. Over time, the plant evolved into S....
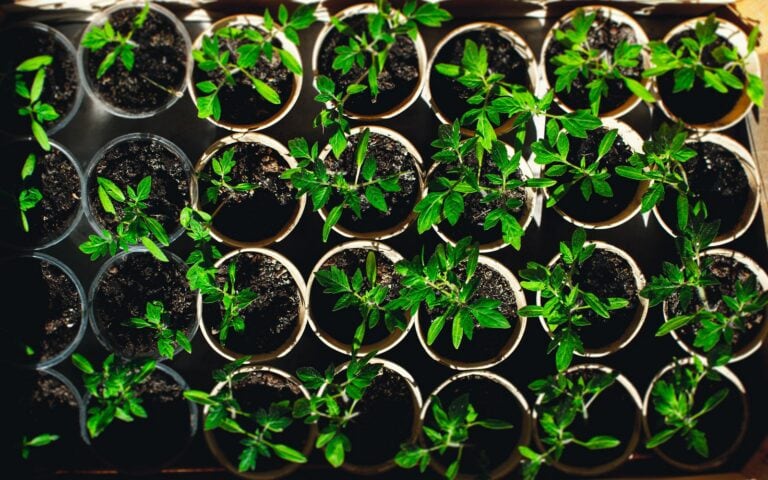

New ‘Tomato Expression Atlas’ dives deep into the fruit’s flesh
From fried green tomatoes to pizza pie, the world savors the tomato across many stages of ripeness, each with its unique qualities. How a fruit ripens has long been an...
Internships
BTI offers a summer research experience program for undergraduate and high school students.
Intern Projects in the Catalá Lab
Investigating the molecular mechanisms underlying fruit set and development
Fruit development is a crucial process in the sexual reproduction of flowering plants and of critical importance for seed dispersal, plant fitness, and agricultural yield. Fruit are complex organs which arise from the coordinated growth and development of floral tissues following pollination. Research in the Catala lab focuses on the molecular regulation of fruit formation and early development using tomato as a model system. We use molecular and genetic techniques to investigate the complex interplay of gene expression changes, signaling events, and hormonal activity, controlling fruit development. The lab also studies the effect of drought stress, an increasing problem in crop production, on tomato fruit set and growth. We are taking advantage of the genetic diversity of wild tomato species to examine the molecular basis of adaptations to water stress and other fruit quality traits.
Previous Interns

Madison Plunkert
Elucidating the function of a novel cysteine-rich peptide during flower-to-fruit transition in tomato
During fruit initiation, secreted peptides mediate communication between pollen and the female gametophyte, as well as the embryo sac and the surrounding female tissues. The cysteine-rich peptides (CRPs), a diverse class of secreted peptides that contain an N-terminal secretion signal and a C-terminal cysteine-rich region, play an important role in cell-cell signaling during flower-to-fruit transition. However, their role in tomato fertilization, fruit initiation, and seed formation is still unknown. Previous tissue-specific transcriptomic studies in our lab identified OVULE-SECRETED PROTEIN (OSP), a CRP highly expressed in the tomato ovule integument at anthesis. Based on this expression profile, we hypothesized that OSP may be involved in processes during flower-to-fruit transition, such as micropylar pollen tube guidance, ovule maturation, or seed formation. In this project, we conducted studies aimed at the functional characterization of OSP, which included phylogenetic analysis, a study of OSP spatiotemporal expression patterns, and the phenotypic characterization of OSP-RNAi plants. In addition, we designed and created molecular tools to generate osp knock-out mutants using CRISPR technology. Multiple sequence alignment and phylogeny tools demonstrated that the cysteine motif in OSP is unique to the cultivated tomato and its wild relatives. Using gOSP::YFP transgenic plants and confocal microscopy, we have observed that OSP is expressed in ovaries and fruit at stages surrounding anthesis. Our results suggest that OSP expression in the ovule inner integument beginning prior to anthesis. OSP-RNAi plants with reduced levels of OSP expression showed no abnormal phenotype when we examined seed weight, seed number per fruit, external seed and ovule morphology, and pollen tube guidance. We have therefore assembled a CRISPR vector that includes guide RNAs targeting the OSP coding region. We anticipate that osp knockout mutants will show a stronger phenotype that will help elucidate the function of OSP.
My Experience
The Plant Genome Research Program at BTI has been an invaluable experience. I felt immediately welcomed by my lab and the community at BTI. In addition to learning a variety of microscopy and molecular biology techniques, I had the opportunity to make experimental decisions and develop confidence as a researcher. Through working independently and with my mentor, I am better prepared for a research career. This internship has reinforced my intention to pursue plant science during graduate school, and I look forward to a continued relationship with my lab and the interns I have met this summer.
Intern Info

Allyson Weir
“Functional Characterization of the Tomato Sugar Transporter SWEET10”
Project Summary:
Carbohydrates produced from photosynthesis are crucial to development, growth, and signaling in plants. To maintain optimal plant function, sugars must be moved from the leaf to other areas of the plant by transporters. A novel class of sugar transporters, named Sugar Will Eventually be Exported Transporters or SWEETs has been shown to play a key role in sugar efflux during several physiological processes, however little is known about their role in fruit development. It has been hypothesized that the action of SWEET transporters is needed in specific fruit tissues and stages, allowing efficient utilization of sucrose during fruit and seed development. However, direct molecular genetic evidence of this hypothesis is still lacking. My project aims to elucidate the role of the tomato SWEET transporter, SlSWEET10, in regulating sugar transport during early fruit development. SlSWEET10 is specifically expressed during early fruit development, and mostly in vascular rich tissues such as the columella and the placenta. The SWEETs gene family is split into four clades, with SlSWEET10 belonging to Clade III. In Arabidopsis, several Clade III SWEETs are known to transport sucrose, therefore we hypothesize that SlSWEET10 is involved in the transport of sucrose from the sieve elements into the fruit parenchyma apoplast. In this work, we have analyzed the effect of variable SWEET10 expression on sugar accumulation and fruit weight using transgenic tomato plants. We have identified the subcellular localization of the SWEET10 protein in fruit, and have compared the expression of SWEET10 and its closest homologs in domesticated tomatoes to their wild relatives. Our results suggest that SlSWEET10 plays an important role in sugar transport during early fruit development, a novel, so far unreported function for SWEET transporters. This research will generate insights into the regulation of fruit development and help identify important genes contributing to crop quality.
My Experience:
Through my time at BTI I have become a more confident researcher. Working independently improved my problem-solving skills and critical thinking, while working together with my lab towards one common project has helped to refine my skills in communication and time management. I felt like I was a member of a wider community of plant biologists whose enthusiasm for their work is infectious and inspiring. Through these experiences I have been able to better define my own research interests and have also been able to speak with Cornell professors about the research being conducted in these areas. I have learned more about life as a graduate student, and the process of applying to Cornell PhD programs. Overall, I believe that this summer has been one of the most transformative and educational in my academic career, and I now look even more forward to pursuing a career in plant biology.
Intern Info

Philip Engelgau
Functional characterization of the tomato sugar transporter SWEET10
Project Summary
Tomatoes are one of the most important fleshy fruit crops of modern agriculture. The sugary globe’s versatile nature has led it to become a staple in the dishes of countless cultures across the world. It is this sugar content that differentiates a refreshingly sweet tomato from an unpalatably bitter one. Although a new family of sucrose transporters called SWEETs has been recently discovered in Arabidopsis and rice, little is known of their actions with regards to sugar acquisition in fleshy fruits in general and tomato fruit in particular. Preliminary RNA-seq data has shown that SWEET10, a member of the SWEET family, is highly expressed during early tomato fruit development in the placenta. Due to this early insight, SWEET10 was selected as a suitable candidate for preliminary functional characterization. Comparison of fruit sugar content in wild-type plants and in transgenic RNAi plants in which SWEET10 expression is down-regulated revealed an altered sugar accumulation pattern in the SWEET10 RNAi lines. Using quantitative RT-PCR we showed that auxin regulates the expression of SWEET10 and some of its closest homologs, a finding which may shed light on its expression, and on the regulation of fruit sugar content during ripening.
Our results not only aim to characterize SWEET genes in tomato, but may also provide insight for breeders hoping to harness genetic information to restore the sweeter characteristics of tomatoes that were lost during domestication.
My Experience
My experiences in the Plant Genome Research Program at BTI will be tremendously helpful in my future scientific endeavors. Not only will the laboratory techniques and skills learned be of great assistance, but the very experience of having spent time at a world-class plant research institute will prove valuable. By observing other researchers collaborate between labs and even between institutes I have gained an understanding of the way science is accomplished. The program has also exposed me to the entire spectrum of plant biology research. Having been shown to such a wide array of the dynamic field of plant biology I feel as though I can make more informed decisions for graduate school and beyond. It is due to the aforementioned traits that my summer at BTI has impressed upon me lessons which will aid me at every step of my future.


