Maria Harrison
Professor, William H. Crocker Distinguished Scientist
Investigating the mutually beneficial partnerships between plant roots and soil fungi that help plants absorb essential nutrients like phosphorus, which is critical for growth.
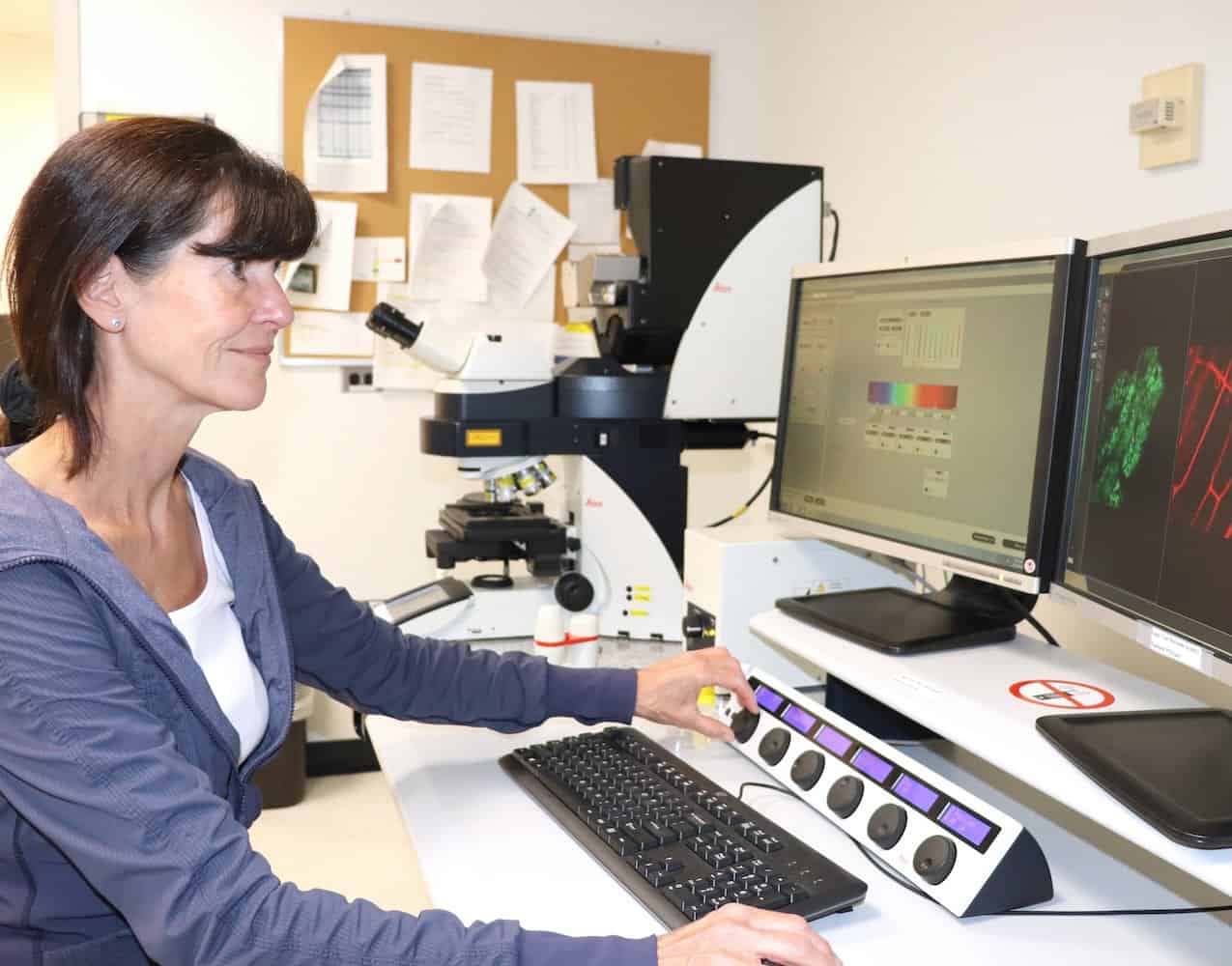

How do plants form symbiotic associations with fungi to access nutrients from the soil?
Email: mjh78@cornell.edu
Adjunct Professor
Section of Plant Pathology & Plant-Microbe Biology
School of Integrative Plant Science
Cornell University
Member
US National Academy of Sciences
Fellow
Royal Society
- Ivanov, S., Müller, L.M., Lefèvre, F.M., and Harrison, M.J. (2026). Yeast two-hybrid-sequencing and bifluorescence complementation resources for assessing protein–protein interactions in arbuscular mycorrhizal roots: CKL2 as a case study. New Phytol. 249, 1592-1604.
- Ivanov, S., and Harrison, M.J. (2024). Receptor-associated kinases control the lipid provisioning program in plant-fungal symbiosis. Science 383, 443-448.
- Lindsay, P.L., Ivanov, S., Pumplin, N., Zhang, X., and Harrison, M.J. (2022). Distinct ankyrin repeat subdomains control VAPYRIN locations and intracellular accommodation functions during arbuscular mycorrhizal symbiosis. Nature Communications 13, 5228.
- Zhang, S.Q., Daniels, D.A., Ivanov, S., Jurgensen, L., Muller, L.M., Versaw, W.K., and Harrison, M.J. (2022). A genetically encoded biosensor reveals spatiotemporal variation in cellular phosphate content in Brachypodium distachyon mycorrhizal roots. New Phytol. 234, 1817-1831.
- Muller, L.M., Flokova, K., Schnabel, E., Sun, X.P., Fei, Z.J., Frugoli, J., Bouwmeester, H.J., and Harrison, M.J. (2019). A CLE-SUNN module regulates strigolactone content and fungal colonization in arbuscular mycorrhiza. Nature Plants 5, 933-939.
- Ivanov, S., and Harrison, M.J. (2019). Accumulation of phosphoinositides in distinct regions of the periarbuscular membrane. New Phytol. 221, 2213-2227.
- Bravo, A., Brands, M., Wewer, V., Doermann, P., and Harrison M, J. (2017). Arbuscular mycorrhiza-specific enzymes FatM and RAM2 fine tune lipid biosynthesis to promote development of arbuscular mycorrhiza. New Phytol. 214, 1631-1645.
- Bravo, A., York, T., Pumplin, N., Mueller, L.A., and Harrison, M.J. (2016). Genes conserved for arbuscular mycorrhizal symbiosis identified through phylogenomics. Nature Plants 2, 1-6.
- Breuillin-Sessoms, F., Floss, D.S., Gomez, S.K., Pumplin, N., Ding, Y., Levesque-Tremblay, V., Noar, R.D., Daniels, D.A., Bravo, A., Eaglesham, J.B., et al. (2015). Suppression of Arbuscule Degeneration in Medicago truncatula phosphate transporter4 Mutants Is Dependent on the Ammonium Transporter 2 Family Protein AMT2;3. Plant Cell 27, 1352-1366.
- Floss, D.S., Levesque-Tremblay, V., Park, H., and Harrison M, J. (2016). DELLA proteins regulate expression of a subset of AM symbiosis-induced genes in Medicago truncatula. Plant Signaling and Behaviour 11, e1162369.
- Zhang, X.C., Pumplin, N., Ivanov, S., and Harrison, M.J. (2015). EXO70I Is required for development of a sub-domain of the periarbuscular membrane during arbuscular mycorrhizal symbiosis. Current Biology 25, 2189-2195.
- Pumplin, N., Zhang, X., Noar, R.D., and Harrison, M.J. (2012). Polar localization of a symbiosis-specific phosphate transporter is mediated by a transient reorientation of secretion. Proceedings of the National Academy of Sciences 109, E665-E672.
- Pumplin, N., Mondo, S.J., Topp, S., Starker, C.G., Gantt, J.S., and Harrison, M.J. (2010). Medicago truncatula Vapyrin is a novel protein required for arbuscular mycorrhizal symbiosis. Plant J. 61, 482-494.
- Javot, H., Penmetsa, R.V., Terzaghi, N., Cook, D.R., and Harrison, M.J. (2007). A Medicago truncatula phosphate transporter indispensable for the arbuscular mycorrhizal symbiosis. Proceedings of the National Academy of Sciences, USA 104, 1720-1725.
- Harrison, M.J., Dewbre, G.R., and Liu, J. (2002). A phosphate transporter from Medicago truncatula involved in the acquisition of phosphate released by arbuscular mycorrhizal fungi. Plant Cell 14, 2413-2429.
- Harrison, M.J., and van Buuren, M.L. (1995). A phosphate transporter from the mycorrhizal fungus Glomus versiforme. Nature 378, 626-629.
Research Overview
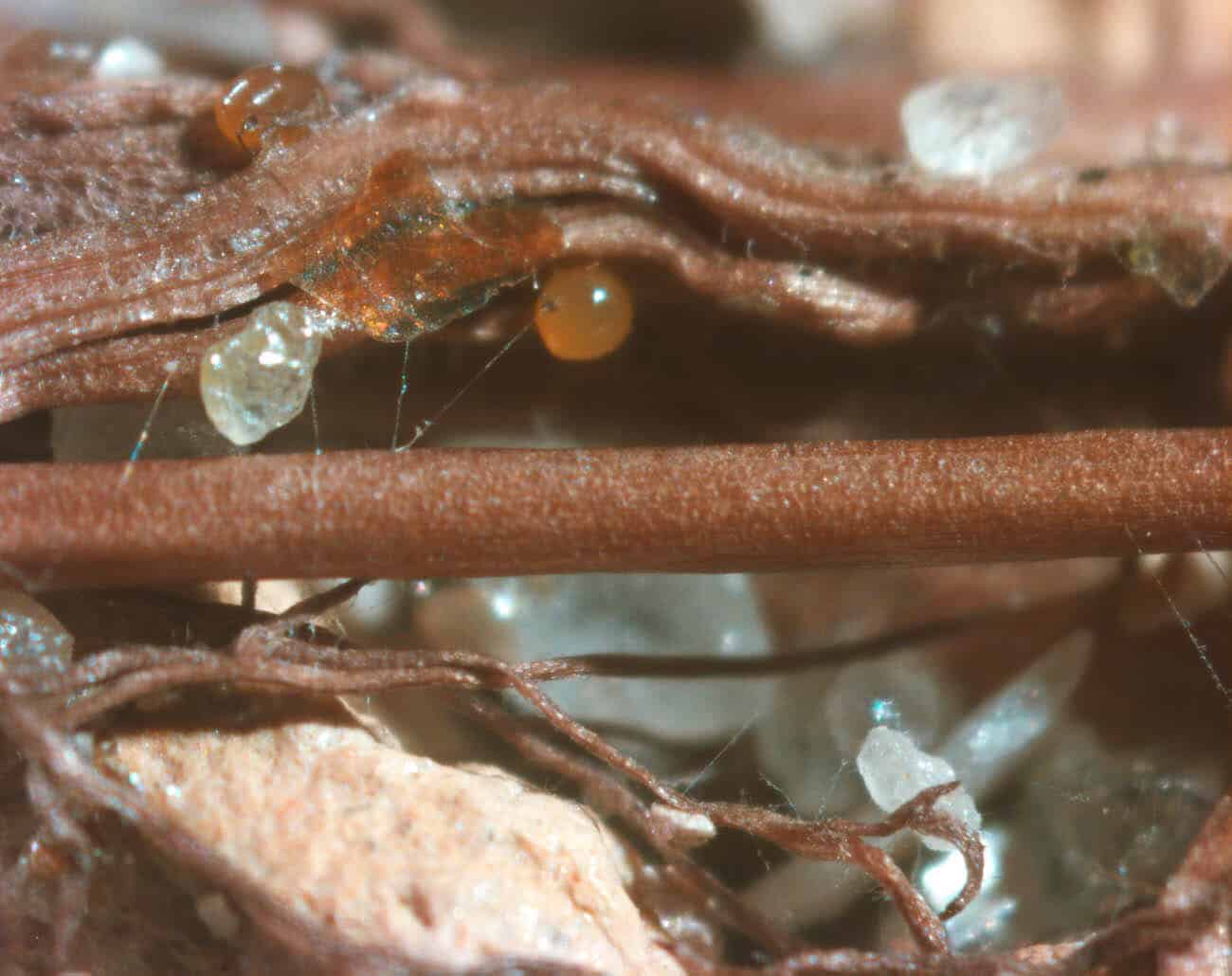
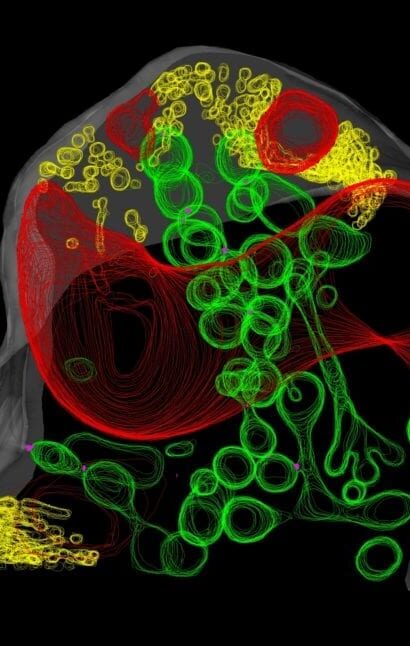
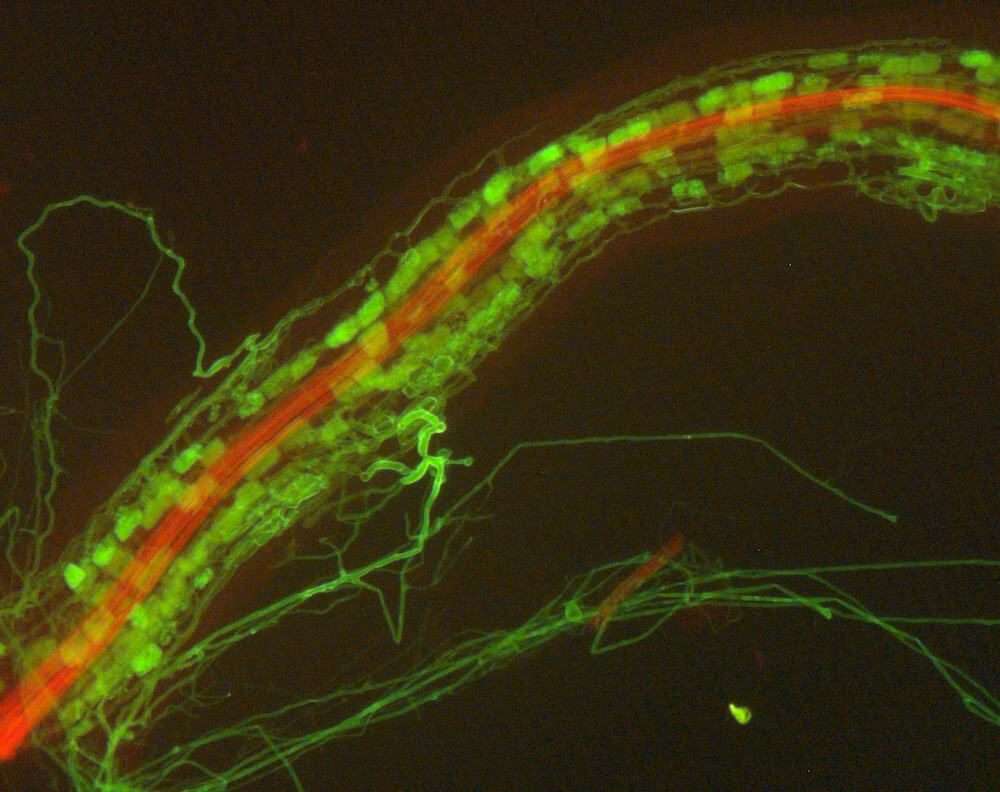
Most vascular flowering plants are able to form symbiotic associations with arbuscular mycorrhizal (AM) fungi. These associations, named ‘arbuscular mycorrhizas’, develop in the roots, where the fungus colonizes the cortex to access carbon supplied by the plant. The fungal contribution to the symbiosis includes the transfer of mineral nutrients, particularly phosphorus, from the soil to the plant. In many soils, phosphate exists at levels that are limiting for plant growth. Consequently, additional phosphate supplied via AM fungi can have a significant impact on plant growth, development and resilience to stress.
The long-term goals of our research are to understand the mechanisms underlying the development of the AM symbiosis and nutrient transfer between the symbionts. Our model organisms include a legume, Medicago truncatula, and arbuscular mycorrhizal fungi, Diversispora epigaea, Rhizophagus irregularis, and Gigaspora gigantea. We use molecular biology, cell biology, genetic, and genomics approaches to obtain insights into the development of the symbiosis, communication between the plant and fungal symbionts, and symbiotic phosphate transport.


Lab Members

Anne Bennion

Lena Cuevas

Natalie Hoffman

Ting-Yu Kuo

Elizabeth Trost

Junjie Wang

Alex Guyon
Postdoc
In the News

Dr. Natalie Hoffmann, recipient of the inaugural Jane Silverthorne Postdoctoral Fellowship at the Boyce Thompson Institute, is investigating how beneficial fungi penetrate plant cell walls without causing damage. Her research explores how plants remodel their cell walls to facilitate symbiotic partnerships, utilizing advanced imaging and genetic tools to solve a mystery with implications for sustainable agriculture.
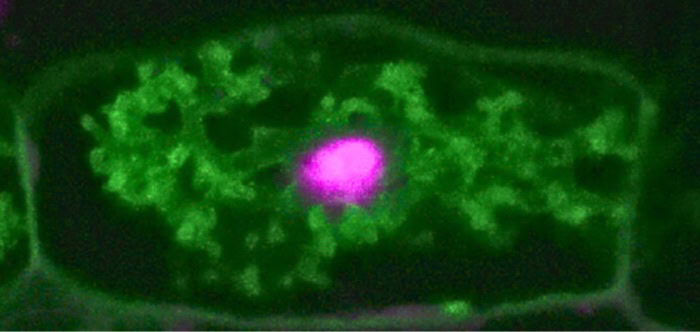
For 450 million years, plants and soil fungi have been trading partners. The fungi weave through plant roots, delivering phosphorus and other soil minerals in exchange for sugars and fats...

Boyce Thompson Institute Celebrates Professor Maria Harrison’s Election to the Royal Society
The Boyce Thompson Institute (BTI) proudly announces that Maria Harrison, the William H. Crocker Professor at BTI and an Adjunct Professor at Cornell University’s School of Plant Sciences, has been elected...
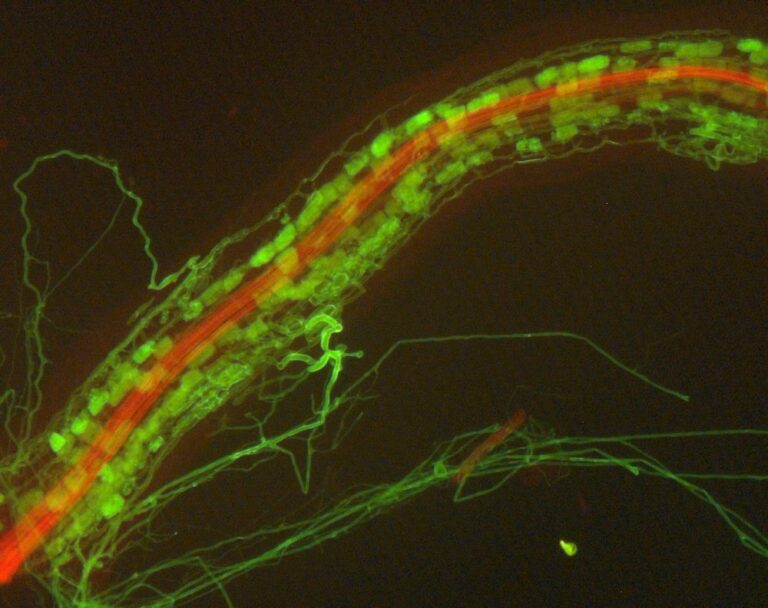
The intricate dance of nature often unfolds in mysterious ways, hidden from the naked eye. At the heart of this enigmatic tango lies a vital partnership: the symbiosis between plants...

Shiqi Zhang spent many months sitting alone in a dark room, staring intently into the lens of a confocal microscope as she focused a laser beam on plant cells mounted...
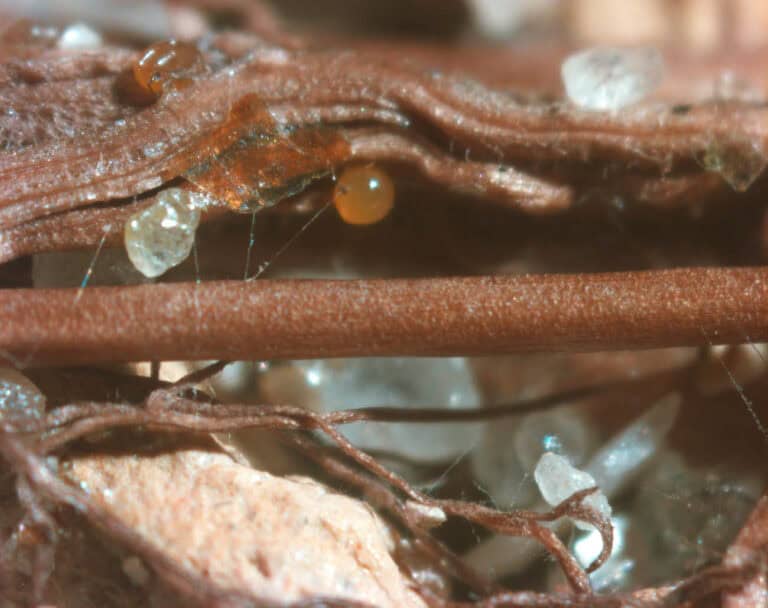

A team of researchers from the Boyce Thompson Institute (BTI) has discovered a distinct group of bacteria that may help fungi and plants acquire soil nutrients. The findings could point...
Internships
BTI offers a summer research experience program for undergraduate and high school students.
Intern Projects in the Harrison Lab
Phosphorus is a critical macronutrient for proper plant growth. While phosphorus deficiencies can be improved by the application of phosphate fertilizers, it is costly, both to the farmer and to the environment. Furthermore, the crops only take up a small percentage of the applied fertilizer; the remainder is either immobilized in the soil, or carried into ground water and rivers, often resulting in pollution.
Interns in the Harrison lab investigate two aspects of plant phosphorus nutrition. The first aspect seeks to understand the basis for the symbiotic relationships between vascular flowering plants and arbuscular mycorrhizal (AM) fungi. The fungi colonize root cells, gaining access to carbon supplied by the plant, while at the same time mobilizing mineral nutrients from the soil, including phosphorus, to be used by the plant. For this work, the lab uses the model legume, Medicago truncatula and the fungus Glomus versiforme. The Harrison lab also studies how plants find and take up phosphorus from the soil when they do not have these symbiotic relationships with fungi. This work toward understanding the mechanisms of perception and acquisition of phosphorus by plants may eventually lead to a more effective usage of fertilizers.
Previous Interns

Olivia Walker
Investigating the role of KINC in arbuscular mycorrhizal symbiosis
Arbuscular mycorrhizal (AM) symbiosis is an over 400-million-year-old relationship where the AM fungus obtains carbon from the plant and in turn provides the plant with inorganic phosphate. Genetically altering plants to better benefit from its fungal partners may mitigate the unsustainable mining of finite phosphate reserves used for fertilizer. The KINASE Cs (KINC1 and KINC2) are a pair of AM host-conserved genes that are essential for AM symbiotic development. Our project investigated mechanism by which the KINCs support fungal accommodation by identifying and evaluating potential KINC-interacting proteins. Bimolecular fluorescence complementation assays were used to evaluate the putative interaction between the KINCs and KIN3. Our data suggest a KINC2 and KIN3 interaction but not a KINC1 and KIN3 interaction. KIN3 was also cloned into our yeast-two-hybrid system for future assays to further access these interactions. Finally, three independent experiments of plants were transformed and planted for tissue in future TurboID proximity labeling proteomics experiments. We additionally aimed to evaluate the importance of KINC1’s kinase activity. This was done by complementation experiments using various single amino acid mutants of the KINC1’s kinase domain, where colonization level was assessed. Our data show that none of the constructs successfully complement the kinc1kinc2 mutant phenotype. However, colonization level was notably low across all root samples. While these experiments must be repeated for confidence conclusions, they represent a first look at mechanisms by which the KINCs function in Medicago.
My time in the Harrison lab was a fruitful experience. My mentor taught me many valuable skills for this field, including Gateway Cloning, Agrobacterium-mediated transformations, confocal microscopy, and more. I am glad to have had the opportunity to work in a molecular lab for the first time, and I look forward to applying the knowledge and skills gained from this experience in future work.
Intern Info

Amitha Karuppiah
Cell-type marker gene expression in Medicago truncatula
Arbuscular mycorrhizal (AM) fungi participate in a mutualistic endosymbiosis with most land plants, exchanging mineral nutrients for carbon compounds through structures called arbuscules. Decoding this symbiosis can contribute to improvements in fertilization, phosphorus conservation, and plant growth. One approach being adapted to AM symbiosis research is single-cell transcriptomics, where RNA-seq data from cells are clustered into cell types using cell-type marker genes. Gene expression in roots, such as in response to AM colonization, can be analyzed by cell type using this method. This project aims to validate putative cell-type marker genes in Medicago truncatula, a model legume for AM symbiosis. Hairy root transformation, GUS staining, and optical microscopy were used to determine if cell-type marker gene expression occurred in the expected cell types. Expression was evaluated in AM-colonized/uncolonized plants (for a putative AM-colonized cortex marker) and across multiple genotypes (wild type R108, Tnt1 mutant della1,2-wtseg [wild type segregant], and Tnt1 mutant GRAS transcription factor knockout lines rad1 and della1,2). Cell-type marker gene expression generally aligned with the expected cell types, aside from two markers where expression primarily occurred in the vasculature instead of the cortex. Expression patterns were similar across genotypes, but with downregulation in the rad1 and della1,2 mutants. Additionally, a root length colonization assay was conducted for the roots transformed with the putative AM-colonized cortex marker promoter::GUS construct. The rad1 and della1,2 mutants showed lower AM colonization than the wild type plants, consistent with the downregulated expression of the AM-colonized cortex marker.
Through this program, I had the opportunity to explore the field of host-microbe interactions by studying a key plant-fungal symbiosis. I enjoyed my research very much, and am grateful to have worked in a supportive environment at the Harrison lab with Trevor Tivey as my mentor. I also became familiar with a variety of techniques, including plant transformation, histological methods, and colonization assays. Outside of the lab, I participated in insightful discussions with graduate admissions personnel, faculty, research/social mentors, and other REU students. These conversations helped me define my goals for the coming years and gain a more nuanced perspective of graduate school and career paths. I liked the variety of events built into the program as well, such as the seminar series and the science communication activity. I will be sure to draw on my experiences from this program at my home institution and in my future research.
Intern Info

Briana Hashim
Investigating a pair of plasma membrane kinases and their impact on arbuscular mycorrhizal symbiosis
Arbuscular mycorrhiza (AM) symbiosis is a mutualistic relationship between plants and fungi supported by over 80% of land plants, including many crop species, making it an ecologically, evolutionarily, and agriculturally important plant-microbe relationship. This association is characterized by intraradical growth of the fungus culminating in the transfer of phosphates and nitrogen to the plant, and in turn, carbohydrates and lipids to the fungi. Therefore, AM symbiosis presents a promising and ecologically sound alternative to reliance on chemical fertilizers to address phosphorus-deficiency in crop cultivation. Despite AM symbiosis’ importance and agricultural utility, little is understood about how host plants are able to undergo the great physiological changes needed to accommodate their intracellular fungal symbiont. To address this question, a phylogenetic study was previously performed to identify genes that exhibit high conservation in AM hosts. In this study, two of these conserved genes that are predicted to encode a homologous pair of leucine-rich repeat receptor-like kinases were selected for characterization. For this purpose, we carried out an extensive array of mutant phenotypic analyses, and utilized RT-qPCR, and prepared for a split ubiquitin yeast-2-hybrid assay, to shed light on the potential roles of these genes in AM symbiosis.
During my time at the Boyce Thompson Institute (BTI) REU program, I had the invaluable opportunity to acquire laboratory skills not readily available at my home institution. A highlight was utilizing a confocal microscope for the first time to image infection units in roots, a fascinating and enriching experience. Interacting with my principal investigator (PI) and mentor was just the beginning; I also established connections with other PIs, mentors, and graduate students from Cornell. This provided me with a valuable insight into Cornell’s graduate student program and the diverse research activities underway at BTI. As a result, my network expanded, and I developed greater confidence in my capability to do scientific research, deepening my understanding of plant-fungal symbiotic relationships. The REU experience has undoubtedly left a lasting impact on my scientific journey, enriching my skills and fostering a sense of collaboration within the scientific community.
Intern Info

Alexis Chun
“Optimizing a deep-learning based program that quantifies arbuscular mycorrhizal colonization”
Project summary:
The arbuscular mycorrhizal (AM) symbiosis is one of the most ubiquitous and important plant-fungal symbiotic relationships. During this interaction, plants are colonized by AM fungi and exchange carbon for otherwise inaccessible essential nutrients, such as phosphorus and nitrogen. Therefore, it is widely studied for its potential to increase nutrient use efficiency in agricultural settings. Fungal colonization is an important parameter for characterizing genes that underlie the mechanisms of the symbiosis, but currently, the most common method for quantifying fungal colonization involves manually counting colonized regions of ink-stained or antibody-labeled roots. This process is not only time-consuming, but also subjective and largely prone to human error. Recently, a deep-learning based program (AMFinder) was developed to automate the quantification process. The program utilizes convolutional neural networks to categorize roots segments in an image as either colonized or noncolonized, providing a much more efficient and consistent method for measuring colonization. The goal of my project was to optimize this program for use in the Harrison lab. To accomplish this goal, the root staining procedure and the imaging method were optimized. The images taken using the optimized protocol were used to retrain the program for a better accuracy. After training with images of the ink-stained roots taken on a flat-bed scanner, the accuracy of the program increased by 5% to 74% accuracy. The small increase is likely due to the inadequate size of the dataset that was used, so in the future, retraining with larger datasets is necessary to reach a higher accuracy.
My Experience:
During my internship with BTI, I had an opportunity to meet and connect with so many fantastic people and was exposed to new topics in plant research. I had never worked with any bioinformatics software previously and had no background in computer science, so this summer was a huge learning experience for me. From working on a machine-learning program for my project to the bioinformatics course, I learned so much about bioinformatics and developed a new set of highly applicable skills. This summer also provided me with an opportunity to learn about graduate school and the insight I gained into graduate programs both at Cornell and in general really helped me clarify my plans for after I complete my undergraduate studies.
Intern Info

Alicia McElwee
“Environmental Impacts on Cytosolic Phosphate Levels in B. distachyon Root Cells During Arbuscular Mycorrhizal (AM) Symbiosis”
Project Summary:
Arbuscular mycorrhizal (AM) fungi form symbiotic associations with the roots of a majority of vascular plants. Their ability to provide phosphate otherwise not accessible to plants through arbuscules has become of great interest in work to optimize the use of non-renewable phosphorus fertilizers. Little is known about how environmental factors impact host plant acquisition of phosphate during AM symbiosis. To assess some environmental factors’ impacts on host root cytosolic phosphate levels, we measured cytosolic phosphate levels across different fungal species and nitrogen conditions using a set of FRET-based phosphate sensors in live roots.
Transgenetic Brachypodium distachyon plants expressing the sensors were 1) colonized with either Rhizophagus irregularis of Diversispora epigaea fungi for testing fungal impact, or 2) colonized with D. epigaea or no fungi and given high nitrogen treatment for testing nitrogen’s impact in AM symbiosis and non-AM symbiosis conditions. The sensor’s signal was measured by imaging the roots’ fluorescence using a confocal microscope. The results showed a difference in root relative cytosolic phosphate levels between the two fungi conditions as well as an increase in root relative cytosolic phosphate levels in the high nitrogen treatment for the non-AM symbiosis condition. No data was collected for the nitrogen AM symbiosis condition due to lack of fungal colonization.
Overall, the results indicate for the first time that relative cytosolic phosphate levels in host roots differ between different fungal species associations and increases with nitrogen application. This emphasizes the additional considerations of environmental conditions if AM symbiosis is to be utilized to optimize crop phosphorus acquisition.
My Experience:
My experience at BTI this summer has allowed me to grow as a researcher, not only introducing me to the field of plant biology through conducting my own experiments but also engaging me with the wider research community of others in this field. Under the guidance of my mentor and others in the lab, I gained confidence in my ability to grasp new experimental techniques and utilize them to develop and test novel questions which build upon our current understandings. Further, the mentoring atmosphere and overall setup of the program showed me first-hand the expanse of resources available for networking and collaboration as I pursue my graduate degree and research career beyond that. The community I was immersed in at BTI and Cornell, including the faculty, students, and my fellow interns, has reinforced my desire to pursue plant research as a field where I can flourish.
Intern Info

Katherine Stahlhut
Discovery of novel gene regulating arbuscular mycorrhizal symbiosis
A majority of vascular plants, including most common crop species, are able to establish intracellular interactions with symbiotic arbuscular mycorrhizal (AM) fungi which provide plants the access to additional sources of inorganic phosphate in exchange for carbon compounds at highly branched fungal structures called arbuscules. The molecular mechanisms involved in subsequent control and maintenance of transcriptional activation during AM symbiosis is largely unknown. Comparative genome analysis identified a previously unidentified gene, which we have tentatively named FBK1, encoding an F-box/kelch-repeat (FBK) protein that is believed to be necessary for AM symbiosis due to its conserved nature in plants species that form the symbiosis. FBK proteins regulate cellular processes through ubiquitination and degradation of target proteins, such as transcription factors. We hypothesize that FBK1 regulates the development of AM symbiosis through the interaction and ubiquitination of transcription factors in the host cell nucleus. The gene is localized primarily in the nucleus of host cells. Plants with a mutated FBK1 genes had higher mycorrhizal colonization rate and less mature arbuscules. A yeast-two hybrid screen of a transcription factor library found that FBK1 interacts with only one transcription factor, RAM1, which is an essential regulator of arbuscule development. The protein FBK1 is therefore believed to be involved in the negative regulation of RAM1, degrading the RAM1 protein in the nucleus in order to decrease AM colonization. This control possibly allows for the host plant to adjust the symbiosis to prevent the exchange of fixed carbon with the AM fungi when phosphorus is not a limiting resource in the environment.
My Experience
During my time at BTI this summer, I have grown as a researcher, having gained a multitude of skills both in lab and for engaging with the world outside. Through planning and carrying out my research project with the support of my mentors, I not only learned hands-on skills imperative for experimentation, but also the interpersonal skills, independence and confidence essential for me in the future. The programming provided by BTI gave me a better understanding of the fellowship, networking and collaborative opportunities available to me as I look towards graduate school and my career beyond. These ten weeks surrounded by faculty, students, and fellow interns reinforced for me that plant science is a vibrant community, one in which I will continue to thrive.
Intern Info

Samantha Clark
“Bacterial Communities in the Hyphosphere of Arbuscular Mycorrhizal Fungi”
Project Summary:
Arbuscular mycorrhizal fungi (AMF) form a symbiotic relationship with plants, providing nutrients from soil, such as phosphorus, while receiving carbon in return. Bacterial communities that associate with AMF hyphae are theorized to solubilize nutrients or directly facilitate or inhibit hyphal growth and exploration of soil. To explore the composition and function of bacterial communities associated with AMF hyphae, an experiment was undertaken to generate a culture library of AMF associated bacterial isolates. Brachypodium distachyon colonized by Glomus versiforme were grown in two compartment microcosms that included a plant compartment in a washed and autoclaved sand/gravel media and a hyphal ingrowth core with a sand/soil (3:1 w/w) matrix. Hyphae were collected from both compartments over four harvests between 14 and 44 days from colonization to 1) retrieve clean hyphae and hyphae directly in contact with the soil matrix and 2) determine if the hyphal communities change over time. Serial dilutions were prepared from harvested hyphae and were streaked on non-selective 10% tryptic soy agar and R2A media. Colonies were purified, the 16S rRNA gene amplified and sequenced to determine taxonomic identity. Overall, 210 bacterial isolates were collected that can be used for future research into their ecology and interaction with AMF. This resource can allow for better understanding of AMF interaction with soil bacteria and eventually lead to possible agricultural applications to improve nutrient management of soils.
My Experience:
As a PGRP intern at BTI, I have gained a confidence in laboratory settings and feel prepared to conquer future research within graduate school. I’ve become very proficient in PCR, gel electrophoresis, and 16S rRNA gene sequencing techniques. Maria’s laboratory was very welcoming, and I learned something new from each and every one. Being on Cornell’s campus has given me the opportunity to meet professors within the plant science department, along with the natural resources department, and has allowed me to obtain an in-depth look at their graduate school programs. Off campus, Ithaca is a beautiful place to live for summer, and I have gained a great appreciation for the area.
Intern Info

Brandon Williams
“Examining the transcriptomal regulators of arbuscule development”
Project Summary:
Many soils are low in phosphorus content and require fertilizer use for farming to be successful. Unfortunately, excessive fertilizer use has negative economic and ecological effects – being a finite resource, not remaining very long in soil, and having the potential to run off and harm adjacent environments. We studied arbuscular mycorrhizal symbiosis, an endosymbiosis between the legume Medicago truncatula and Glomeromycota fungi. In this relationship, the plant provides the fungi with carbon in exchange for phosphorus. This association has been shown to positively impact the growth of the plants involved and could potentially lead to the decreased use of fertilizers.
Our project examined one of the major symbiotic transcription factors, IPD3. It was recently found that M. truncatula possesses two copies of the gene, IPD3A and IPD3B. Our work employed the yeast two hybrid approach to explore the two genes’ interactions with other transcription factors and to compare the two with each other. Furthermore, we set up a phosphate, growth response, and root architecture experiment to gain additional insight of the differences between the wildtype and IPD3 mutant variations of M. truncatula. The ultimate goal was to gain a better understanding of the mechanisms that play a role in AM symbiosis so that we can better employ the nutrient-capturing abilities of AM fungi. Further research into AM symbiosis could prove fruitful as the association has the potential to impact ecosystems’ carbon cycle and productivity.
My Experience:
My time here at Boyce Thompson Institute has been extremely rewarding. Getting to know some of the other interns has helped me build connections that I know will last indefinitely. The same can be said about the Harrison lab which was more than welcoming and fostered a family-like community. I was able to draw from everyone in the lab, everybody offering something different, allowing me to build upon my molecular biology and practical skills.
I am more than grateful for my mentor for being so patient and willing to offer her insight whenever needed. Getting to learn the yeast two hybrid system was one of my favorite parts of this experience, knowing that it is a useful technique out in the world of science. Leaving here, I feel more confident in myself and in my decision to pursue a graduate degree. Any undergraduate should take advantage of this program if given the opportunity!
Intern Info

Ethan Thibault
Investigation into Root Colonization of Wild Rice and Brachypodium distachyon by Arbuscular Mycorrhizal Fungi
Project Summary
With an increasingly desperate demand for food and renewable energy, researchers are interested in producing higher yields in agronomic and biofuel plants like rice, wheat, barley, Miscanthus, switchgrass, and Prairie Cordgrass. Arbuscular Mycorrhizal Fungi (AMF) are symbiotic organisms that penetrate the roots of all of these plants and colonize their cortical cells. At these colonization interfaces the fungi provide the plant with nutrients like phosphate and nitrogen from the surrounding soil, while the plant provides the fungus with carbon for its growth. Colonization typically leads to increased tiller production, biomass, and yield. Understanding how AMF enter and move through root cells is crucial to maximizing colonization, which can lead to less fertilizer application, increased yield/biomass in agronomic and biofuel plants, and the ability to grow them on marginal lands. To do this, we are inoculating 17 inbred accession lines of Brachypodium distachyon and 8 lines of wild rice with AMF. Based on the known phenotypic variation within these groups, we expect differences in the amount of colonization and the patterns of infection. Expected changes in gene expression from line to line will lend insight into the mechanisms of colonization as well. We are also creating CRISPR knockouts of multiple candidate cell wall proteins that are upregulated during symbiosis with the fungi. These same genes and their promoters are being tagged with GFP to identify subcellular localization and monitor expression patterns respectively. We expect this to yield insight into how the root cell walls remodel to accommodate the growing fungi.
My Experience
It has been an incredible two months here at BTI. I have learned so many new lab techniques like cloning, bacterial transformations, primer construction and more. I was able to explore the molecular aspects of plant biology in a way that I couldn’t before and I was able to take part in a professional research environment with individuals at different stages of their career. They offered insight into what to expect in graduate school and as a postdoctoral researcher. It was also great being able to explore Cornell University, knowing that I want to apply here for graduate school, and also knowing that I could come to BTI for my graduate degree. Finally, the best part about partaking in this experience, for me, was to be completely surrounded by other researchers who are just as passionate as I am about plant biology and the potential for plant research.
Intern Info

Raphael Amsili
New Visions student, TST-BOCES, September 2014-August 2015
Ithaca High School senior
Raphael contributed to a project studying arbuscular mycorrhizal symbiosis in Medicago truncatula.
Intern Info

Sara LeFevre
Identification of protein interactors in Medicago truncatula exocyst complex
Project Summary
Most vascular land plants participate in a beneficial relationship with arbuscular mycorrhizal (AM) fungi; in which the plants provide carbon to the fungi in exchange for phosphates and nitrogen. This exchange is mediated by a variety of mechanisms within the root and the fungal structures that enter the root cells, known as arbuscules. Arbuscules are surrounded by a plant-derived membrane, the periarbuscular membrane (PAM), which forms an interface where controlled exchange of nutrients takes place. Plant cells build the PAM by redirecting the exocytotic pathway. Vesicle fusion in exocytosis is executed by an octameric protein complex called an exocyst. Research in the Harrison lab showed that a knockout mutant of exo70i, a subunit of the exocyst complex, was unable to support normal arbuscule development (Zhang, et al., 2015). In order to confirm that MtEXO70I is a true subunit of the exocyst complex, split ubiquitin yeast two hybrid mating experiments were performed to visualize protein-protein interaction between EXO70 and other exocyst subunits. This experiment demonstrated that MtEXO70I is a true subunit of the exocyst complex. A second experiment explored the activation of MtEXO70I. Cell polarity and activation of the membrane exocyst complex in yeast and mammals is reliant on the small GTPases of the Rho family. In plants, these are called ROP (Rho of Plants) GTPases. To determine if the same mechanism of activation exists in plants, split ubiquitin yeast two hybrid analysis was performed between ROP GTPases and EXO70. These experiments provided evidence that MtEXO70I is regulated by ROP GTPases. Research that expands our knowledge about the mechanisms which cause exocytosis in plants can lead to a greater ability to control the beneficial symbiosis that occurs between plants and mycorrhizal fungi; for example, subunits of the exocyst complex may be manipulated to cause a longer symbiosis, or cause the plant to uptake a larger quantity of nutrients. This research is important because it will enhance the ability of molecular biologists to manipulate plants in order to improve/develop sustainable agriculture.
My Experience
My summer spent at Boyce Thompson Institute was as challenging as it was rewarding. I truly learned what it means to do science: asking questions, understanding not only how to do a technique but why, using both precision and accuracy, never taking anything at face value, never cutting corners, and not giving up when the first, second, or hundredth time was a failure. The core skillset I am taking away from this experience is not simply the techniques I learned, but the ways to learn new techniques in the future, such as how to read a protocol or glean important information from a dense scientific paper. In addition, I learned how to communicate with other scientists for their expertise on any subject. This was a brief, intense, and truly worthwhile internship. I feel more prepared now than ever to pursue a career in biological sciences.
Intern Info

Carin Ragland
Identifying AM symbiosis genes in Medicago truncatula through reverse genetics
Project Summary
Arbuscular mycorrhiza (AM) is an endosymbiotic association between biotrophic fungi and vascular plants in which the fungus supplies water, phosphorus, and other nutrients to the plant and receives carbon in return. AM is a potentially powerful fertilizing tool in agriculture because, in some cases, fungal interactions increase the amount of nutrients accessible to the plant without increased external nutrient applications to the soil. About 80% of angiosperms can form AM, and it is considered one of the most ubiquitous plant symbioses on earth.
In two experiments during this internship, two gene deactivation methods—RNA interference (RNAi) constructs and insertion lines—were used to test several genes proposed as AM participants by phylogenic and bioinformatic analyses. The RNAi constructs were prepared via molecular cloning and introduced into M. truncatula seedlings by root transformation. The RNAi constructs and insertion lines of the candidate genes were cultivated in “cone-tainers” with 200 fungal spores for three weeks and then harvested. Root samples were collected from the insertion lines and stained with fluorescent dye. RNAi construct samples were selected via a Red Root fluorescent marker before staining. Root colonization and arbuscule development were assessed in the insertion lines and RNAi transgenic roots to determine whether gene deactivation affected AM symbiosis.
Studies featuring gene deactivation methods (reverse genetics) have made significant progress in explaining the molecular mechanism behind AM. With continued AM research future discoveries could generate widespread use of AM symbiosis within agriculture to increase crop yields and address resource conservation concerns.
My Experience
For two years I worked with sustainable agriculture projects, informally utilizing and studying plant science concepts such as microbial and fungal symbiosis within urban agriculture initiatives. I was excited for the opportunity to participate in formal plant science research regarding AM symbiosis because it is such a fundamentally crucial ecological interaction. I enjoyed my experience at BTI, and truly appreciate the time my mentor, PI, and lab mates dedicated in contributing to my first research experience. This summer I have learned widely applicable lab techniques, implemented an independent project, and became acquainted with young scientists from a myriad of different fields. The program demystified the graduate school experience and application process and exposed us to useful resources and knowledgeable researchers and educators. In the future I will definitely continue plant research and am considering graduate studies in Agronomy.
Intern Info

Celina Abundis
Exploring gene function in AM Symbiosis
The purpose of my research this summer was to investigate the potential role of interesting fungal genes in the Arbuscular Mycorrhizal (AM) symbiosis. AM fungi are obligate biotrophs; dependent upon another organism for nutrients. AM symbiosis occurs between vascular flowering plants and Glomeromycota fungi. Little is known about the fungal genes that control AM development. In the Harrison lab I was able to verify the expression of genes in Gigaspora gigantea and Glomus intraradices, AM fungi, with gene-specific primers. I verified the genes are fungal through PCR on their corresponding AM fungal genomic DNA, and then showed their expression with reverse transcriptase (RT) PCR using AM fungi inoculated Medicago truncatula roots. Agrobacterium rhizogenes was used to generate M. truncatula plants with transgenic roots overexpressing the G. intraradices gene under an AM-specific promoter. This was done by making a compatible binary vector via Gateway© cloning system. Seeds were first germinated on a petri plate and transplanted to cones where they are inoculated with G. intraradices fungal spores and harvested after 2-3 weeks. My next step was staining roots for the presence of intraradical colonization and visualizing the AM phenotype with a microscope. In the case that the gene is important for AM symbiosis, I predicted that overexpression would have a positive impact on AM development and promote greater fungal colonization.
My Experience
My experience in the Harrison lab has been beneficial for the next step in my academic career. With the help of my mentor, I’ve learned several new techniques: colony PCR, RNA extraction, RNAi constructs, Agrobacterium root transformation, staining and visualization of AM fungi within roots, and staining for GUS expression. The other lab members also made my experience enjoyable, even when a mistake was made. I plan on applying for graduate schools this fall, and I am interested in programs that specialize in Plant Pathology and Plant Breeding and Genetics.
Intern Info

Alexander Platero
MtPT4: A vital protein for arbuscular mycorrhizal symbiosis within Medicago truncatula
Although the importance of phosphorous in overall plant health is well known, the acquisition of phosphorous from arbuscular mycorrhizal (AM) fungi to plant root cortical cell in AM symbiosis is still not well understood. However, using the model legume, Medicago Truncatula, certain components involved in the mechanism have been illuminated. MtPT4 is a proton coupled phosphate transporter located on the periarbuscular membrane of cortical cells undergoing symbiosis. It has been shown through research that this protein is necessary for not only phosphate transport into the plant, but for the morphological changes that is required to form the symbiotic infrastructure between fungus and plant. From the identification of the MtPT4 gene, transcription factors leading to the expression of this gene have been found. A designated MtTF1 is one of these transcription factors. Its expression has been shown to only occur in cortical cells containing arbuscules of the fungi. It is speculated that TF1 is not alone in the process of expression and using cDNA libraries from gene data banks, 60 proteins have been selected as potential candidates. Using the yeast two hybrid method, several of these proteins have been confirmed as interacting partners of TF1. Now further testing is needed to conclude that the proteins do interact. Another area of study is the expression pattern of MtTF1. Expression of MtTF1 has been shown to be significantly high in roots undergoing colonization and almost no expression exists elsewhere. Thus, the expression pattern of this transcription factor is also of interest and is currently being studied using promoter-Gus staining analysis of the roots of plants undergoing symbiosis with AM fungi.
My Experience
The Harrison lab partakes in unraveling the mystery behind the mechanisms that occur between arbuscular mycorrhizal fungi and plant symbiosis. I was fortunate to be part of that team. The experiences that I had in this lab have been ineffable pleasures. The lab members were tremendously amiable and enamored by their projects. At first, the information seemed insurmountable as everything was new, but the lab members made me feel at home. Soon I began understanding the new lab experiments and techniques used in genomic plant research. Having mentor guidance gave me an ambivalence of excitation and intimidation. I felt excited because I never knew what new technique I was going to learn from my mentor and I felt intimidated because my knowledge of this new universe was inadequate in comparison to his. However, as time progressed, I became much more confident in my abilities as I learned lab techniques and was eventually allowed to conduct work independently (After making a few mistakes of course). The time spent here has definitely solidified my plans of going into the research field. It is an experience I would have all over again.
Intern Info

Lauren Carley
Investigating the role of GA signaling in arbuscular mycorrhiza
Arbuscular mycorrhizal (AM) symbiosis is a mutualistic plant/fungus interaction that can be formed between almost all land plants and fungi belonging to the phylum Glomeromycota. This relationship is important because it provides plants with increased nutrient uptake, especially for limiting nutrients such as phosphorous and nitrogen. As a part of the overall effort in the Harrison lab to elucidate the molecular basis of this symbiosis, my project focused on gibberellin (GA) signaling and its role in the formation of arbuscules, the functional units of AM symbiosis. I characterized the AM phenotypes of several Medicago truncatula lines carrying mutations in genes involved in the GA signaling pathway, thereby helping to understand how GA signaling regulates arbuscule formation. I also transformed plants with constructs that allow for spatial localization of the expression of genes believed to be targets of the GA pathway.
My Experience
During this internship, I learned several new techniques, including Agrobacterium rhizogenes-mediated root transformation, staining and visualization of AM fungi within roots, and histochemical staining for GUS expression. Other tasks included the generation of a promoter-GUS fusion construct, genotyping an M. truncatula mutant line, DNA and RNA isolation, fertilizer preparation, and general plant care. I worked closely with my mentor, post-doctoral fellow Daniela Floss, throughout the summer, and also received help and support from all members of the Harrison lab.
Intern Info

James Eaglesham
Plant-Fungal Symbiosis
During this summer, I worked on the symbiotic interaction between Medicago truncatula and glomeromycotan fungi. My project was focused on the study of nutrient transporters present in root cell colonized by these mycorrhizal fungi. Therefore, I examined several single and double mutant phenotypes by using microscopic techniques and semi quantitative polymerase chain reaction. I found several discrepancies in total arbuscule number and gene expression between the mutants and wild type plants. Taken together, these data revealed the necessary presence of these transporters for a functional symbiosis.
In conclusion, I learned to work with a plant-fungal symbiosis and understood its ecological and possible economical benefits. I learned many different techniques and I really enjoyed my time working at BTI and have continued this project throughout the school year.
My Experience
During this summer, I worked on the symbiotic interaction between Medicago truncatula and glomeromycotan fungi. My project was focused on the study of nutrient transporters present in root cell colonized by these mycorrhizal fungi.
Intern Info

Darrin Schultz
Natural variation in arbuscule development and lifespan in selected ecotypes of Medicago truncatulaarbuscular mycorrhiza (AM) symbiosis with Gigaspora gigantea
My experiments were a part of the larger Medicago HapMap project, an international research consortium that seeks to explore the genetic diversity of the model legume, Medicago truncatula and to identify novel genes involved in various biological processes, including symbiosis. I screened transcript levels of symbiosis-specific genes in 30 ecotypes of M. truncatula when colonized with the arbuscular mycorrhizal (AM) fungus, Gigaspora gigantea in order to understand the variation in AM symbiosis among these ecotypes. My mentor and I found a large amount of diversity in the transcript levels of these genes, thus showing a variation in AM symbiotic status. The data generated from this experiment will be used in future Genome Wide-Association studies including all 384 Medicago HapMap accessions to identify novel genes that play a role in AM symbiosis.
Intern Info

Timothy Heslop
A suppressor screen of Medicago truncatula mtpt4-1 for gene mutations in the regulation of arbuscule degradation
We are screening mutagenized Medicago truncatula mtpt4-1 mutants for suppressors of the mtpt4-1 phenotype. The pt4-1 mutant varies from the wild type in that it speeds up the expiration of arbuscules in root cells. The wild type allows for the natural development and death of arbuscules. The found mutants will latter be used in identifying the genes responsible for arbuscule regulation.
Intern Info

Vanessa Powell
My project involved working with arbuscular mycorrhizal (AM) fungi which forms symbiosis with most vascular plants. The project focused on looking at expression patterns in genes encoding transcription factors.
Intern Info

Julie Grimm
Julie is currently a junior at Whitman College in Washington. During this past summer in the Harrison lab, Julie worked with her mentor Karen Gomez on differential gene expression in Medicago truncatula mutant and wildtype colonized by Gigaspora gigantea. While in the Harrison lab, Julie used common molecular biology techniques such as RT-PCR, cryosectioning, and RNA extraction.
Intern Info

Zachary Frederick
Zachary is currently a senior at student Trumansburg High School in NY. During this past summer in the Harrison lab, Zachary worked with mentor Karen Gomez. Zachary’s project consisted of studying the short-term effects of phosphate on Medicago truncatula wild-type, str and mtp4-1.
Intern Info

Zach King
Zach is currently a senior attending SUNY College of Environmental Science and Forestry. As an intern, Zach worked under the supervision of his mentor, post-doc Julian Levy on the role of Gibberellic Acid related genes of Medicago truncatula and their Involvement in the Arbuscular Mycorrhizal Symbiosis. In the process of completing his project, Zach learned several ‘wet lab’ skills such as reverse transcriptase PCR, gel electrophoresis, and microscopy.
Intern Info

Stephen Mondo
Stephen is currently a student at the SUNY Binghamton. During this past summer in the Harrison lab, Stephen worked with mentor Quan Zhang. Stephen’s project consisted of fine mapping of a mycorrhizal mutant in Medicago trunculata. To do this, Stephen learned several techniques including; enzyme digestion, bacterial transformation, DNA extraction, nad fluorescence microscopy.
Intern Info

Jose Arevalo
Jose is now attending the University of California at Berkeley. As an intern, Jose worked under the supervision of his mentor, post-doc Julian Levy on the silencing of kinases and nodulin transripts imiplicated in arbuscular mycorrhizal symbiosis in Medicago trunculata. In the process of completing his project, Jose learned several ‘wet lab’ skills such as reverse transcriptase PCR, gel electrophoresis, bacterial innoculation, and plasmid DNA extraction.
Intern Info

Cory Ellison
Cory is currently a student at Baldwin-Wallace College. During this past summer in the Harrison lab, Cory worked with mentor Jinyuan Liu. They transformed Medicago truncatula to over-express a serine threonine kinase and found that the mutants couldn’t express these proteins and were unable to form symbiotic associations with arbuscular mycorrhizal (AM) fungi.
Intern Info

Tom Kraft
After spending the spring semester in the Harrison lab part-time through a research-based class at Lansing, Tom returned full-time to continue his project for the summer. Tom worked directly with Maria Harrison to study Medicago truncatula mutants in order to gain useful insight regarding phenotypes of roots, leaves, and the mycorrhizal association. Specifically, he mainly worked with a mutation called “chicken feet”, which occurs in the roots of mutant plants.
When asked about his hobbies, Tom responded emphatically, “I love to unicycle!” He added that unicycling only compliments his clown dreams, which also include juggling and using a Chinese yoyo.




















